Cell Systems and Mechanobiology Lab
About
We create physiologically realistic computational models of cells and organs to advance healthcare. Our "digital-twin" models and physiology-informed algorithms advance basic understanding of how biological systems work and inform decision making in clinical and pharmaceutical applications.
Biology has provided an enormous body of descriptive data in its bid to understand the organism. But these descriptions do not explain the dynamic organisational capacity of cell-based life.
At the Faculty of Engineering and Information Technology, the Cell Systems and Mechanobiology Lab is applying engineering sciences to create new ways to integrate and analyse biological data. We bring advanced imaging technology, biomechanics, systems biology, mechanobiology, mathematical modelling and artificial intelligence together within a vibrant, intellectually stimulating, multi-disciplinary team to solve fundamental mysteries of how biology works. We apply our algorithms and technologies to solve disease treatment, diagnosis and drug discovery applications.
Our goal is the development of dynamic simulations that explain changes to cellular systems, with models providing a new platform to test and develop new disease interventions.
Research Areas
We are experts at creating anatomically accurate and physiology-informed computational models of living cells and organs. In the process of creating these models, we uncover fundamental mechanisms underpinning the function of biological systems in health and disease. Our major application focus is cardiovascular disease, which is the leading cause of death in Australia and the developed world. Our basic-science focus is to advance our understanding of how cells work using innovative computational models and techniques. We list a selection of our current projects below. Current opportunities to join our lab are posted at the link at the bottom of this page.
Cell physiome
Underpinning our biomedical research are engineering innovations we use to create our computational models. We conduct research in AI-based microscopy image segmentation algorithms, mathematical modelling and computational techniques such as finite element modelling and coarse-grained molecular modelling.
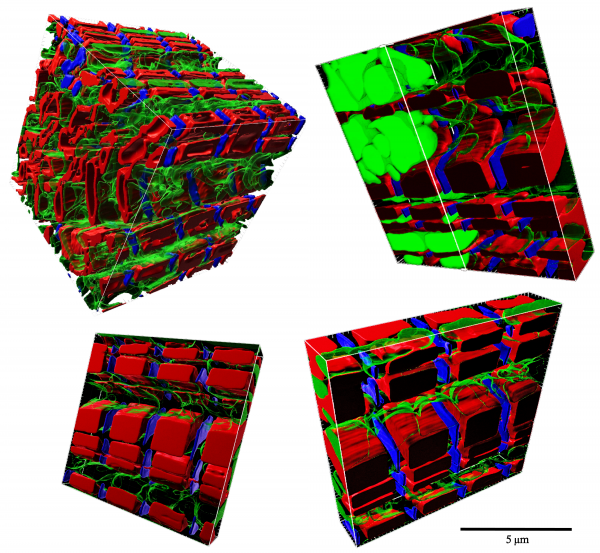

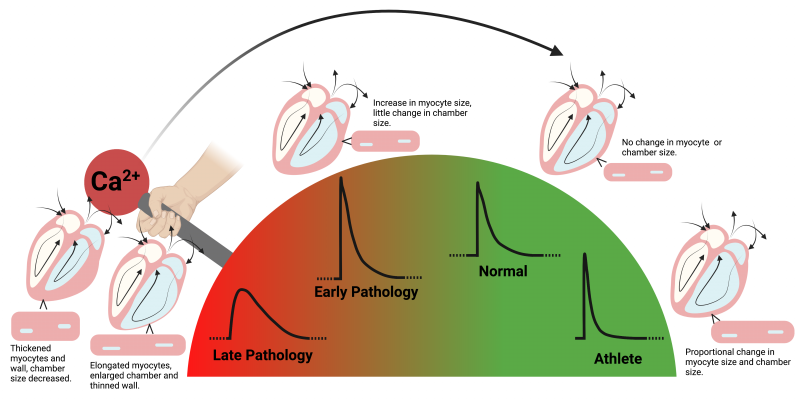
Cardiac hypertrophy
Our hearts grow as a response to changes in blood supply demand across our lifetime. For example, the heart grows from the size of a walnut to the size of our fist from birth into adulthood. The heart also grows as a response to diseases that cause increased blood pressure or demand for blood supply, as in hypertension for example. But this growth often exacerbates the disease. A precise understanding of how heart growth, also called cardiac hypertrophy, is regulated will unlock new therapies and tissue engineering technologies. In this project we are addressing this fundamental gap in our knowledge of heart muscle remodelling by studying the role of mechanics, metabolism and calcium on hypertrophic remodelling.
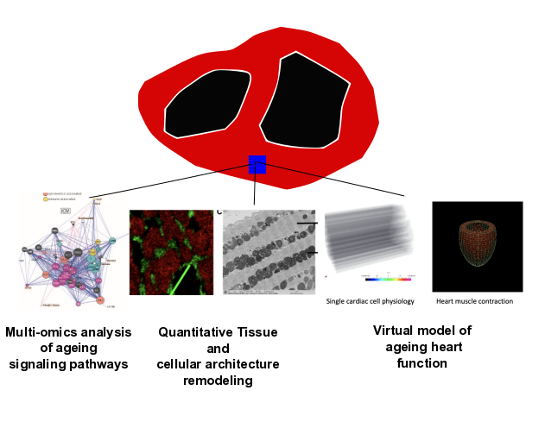
Systems biology of the ageing heart
Why are we at increased risk of heart disease as we age? This is an increasingly important question to answer with an ageing Australian population. We are partners in a cross-institutional and cross-disciplinary research program between Melbourne and University of Sydney to create an atlas of the molecular changes to ageing human heart. We are developing new artificial intelligence algorithms and computational models of the heart that we will use to visualize the structural, molecular and functional changes to the heart as we age.
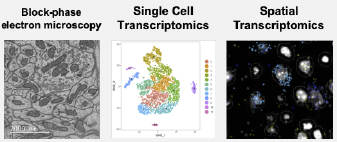
The developing heart at single-cell resolution
Non-compaction cardiomyopathy (NCC) is a congenital heart disease that is considered the third most diagnosed cardiomyopathy in adults. Current consensus points to the process of cardiac compaction and chamber formation during heart development as the origin of NCC. However, very little is known about NCC genetic determinants as most of the ones identified are like other cardiomyopathies. We are applying innovative deep learning segmentation and image analysis algorithms to map out the morphological and genetic changes that govern chamber formation in the developing heart in health and NCC.
Red blood cell and vascular biology
Red blood cells must pass through narrow capillaries half their diameter to supply oxygen to all parts of the body. Changes to their biophysical properties can occur through parasite infection (as in malaria) or genetic causes (as in sickle cell anaemia) and affect their passage through capillaries. Similarly, changes to the cells that make up blood vessels can also affect red blood cell transport. We are collaborating with CSL to develop microfluidics based vasculature-on-chip devices for high throughput drug-screening assays and to understand the fundamentals of red blood cell and vascular interactions.
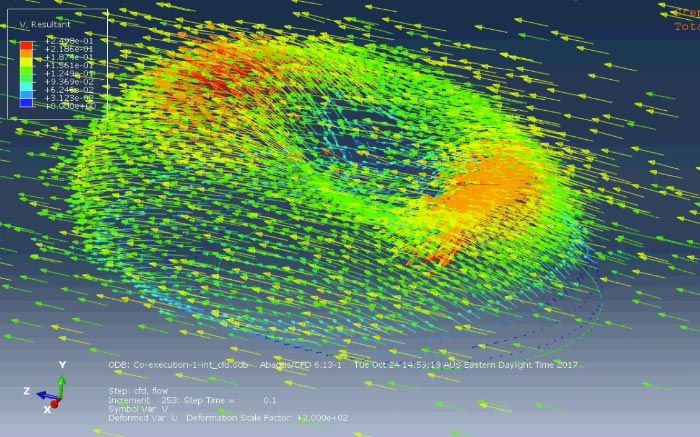
Cell-cell adhesions in tissue formation and immunotherapy
Cells are the building blocks of multi-cellular living systems. The process of cells joining to form tissues and then to maintain them is regulated by an intricate network of chemical reactions and mechanical forces acting at the cell-cell adhesion junction. Cell-cell adhesion disruption by cancer leads to spread of cancer to other parts of the body and immune cells form adhesions with viruses and cancer cells before killing them with toxins. We are creating a new digital-twin model to show how key molecular players work in concert to form cell-cell adhesions.
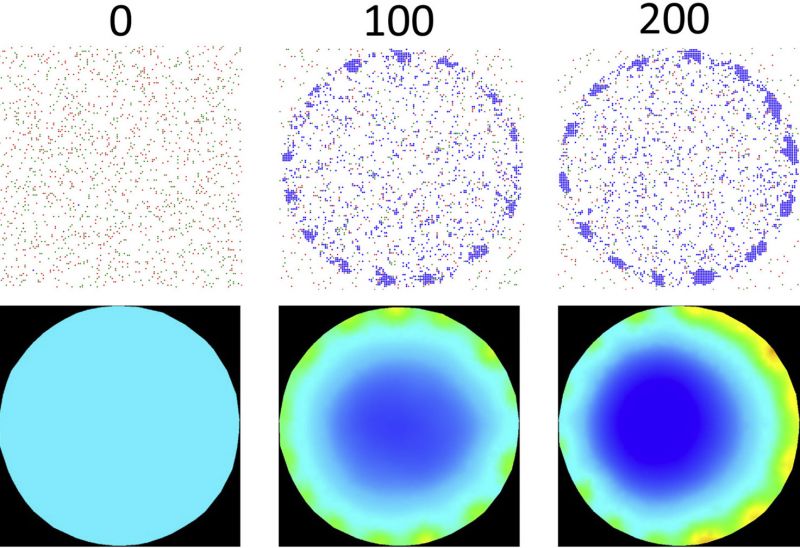
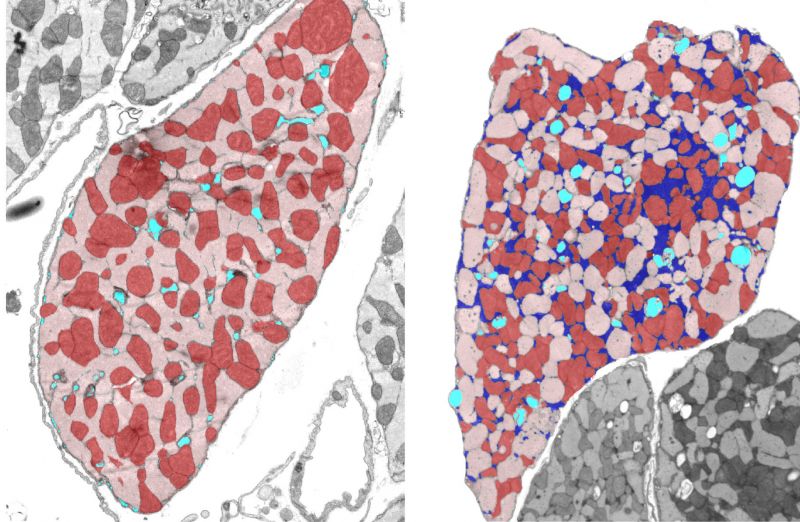
Mitochondria and cellular metabolism
Mitochondria produce the energy needed to sustain cellular life. Proper mitochondria organisation and function is therefore critical for health. Dysfunction and disorganisation of mitochondria within cells are associated with many diseases in the body including parkinsons disease, heart failure and more. But we are yet to understand fundamentally how mitochondria organisation and dynamics regulate energy production and consumption. We are developing mathematical models of mitochondrial bioenergetics and dynamics to address this gap and to discover new strategies to treat disease conditions by targeting mitochondria.
Group leader
A/Prof Vijay Rajagopal
Staff
Graduate researchers

Rachel Alexander
Microfluidics for vasculature-on-chip based drug screening
alexander.r@student.unimelb.edu.au
Meezaan Dickinson
Modelling Ca2+ Signalling in Cardiac Hypertrophy using a Bond Graph Approach
m.dickinson1@student.unimelb.edu.au
Volkan Ozcoban
Mathematical and Computational Modelling of Mitochondrial Dynamics during Cell Migration
vozcoban@student.unimelb.edu.au
Aidan Quinn
Multi-source correlative microscopy image generation with deep-learning
aidan.quinn@student.unimelb.edu.au
Ashan Wanasinghe
Development of a multi-parameter measurement platform for single cell resolution phenotyping of cardiac cells
ashan.wanasinghe@student.unimelb.edu.auAlumni
| Name | PhD topic | Current role |
|---|---|---|
| Giovanni Guglielmi | Cardiac growth and ageing | University of Sydney |
| Afshin Khadangi | Machine learning | Tally Group |
| Jared Collette | Cancer cell motility and mechanobiology | Walter & Eliza Hall Institute (WEHI) |
| Qilin Yu | Cancer cell motility and mechanobiology | neoX Biotech |
| Arman Namvar | Red blood cell mechanics and malaria | CSIRO |
| David Ladd | Modelling cellular dynamics | oNko-Innate Pty Ltd |
| Agne Tilunaite | Cell Signalling | InterAx Biotech |
| Hilary Hunt | Cell signalling | University of Oxford |
| Shouryadipta Ghosh | Diabetic cardiomyopathy | CSIRO |
| Maciej Kubicki | Breast density and breast cancer risk | |
| Jan Jarosz | Diabetic cardiomyopathy |
Contact us
- Address
- Building 261, 203 Bouverie Street
Department of Biomedical Engineering
The University of Melbourne
VIC 3010 Australia - vijay.rajagopal@unimelb.edu.au
- Phone
- +61 3 8344 1281
- Twitter: Group: @cellsmb

